Proteins are the building blocks of living cells and tissues. So when these biomolecules “misfold”—fail to self-assemble into their functional, three-dimensional forms—or malfunction in some other way, it can cause all sorts of health issues, including neurodegenerative disorders (such as Parkinson’s disease and Alzheimer’s) and even cancer.
Jeffrey Weber, the winner of the Hertz Foundation’s Thesis Prize for 2015, has taken a novel approach to understanding the dynamics of protein folding, blending ideas from statistical physics, biology and chemistry to find out how proteins function, why they get trapped in misfolded states, and how they connect to energy sources and the living environment within cells.
“Life is fundamentally a nonequilibrium phenomenon, so studying proteins in space-time matters,” Weber said. “If some dynamical process is harmful or toxic on timescales relevant to individual cells, that’s of course a problem for humans.”
Weber’s doctoral thesis, “Far-From-Equilibrium Phenomena in Protein Dynamics,” is a culmination of his research at Stanford University. Weber utilized molecular dynamics simulations, complex network models, and nonequilibrium theories to study protein molecules at an atomic level. Using data derived from the Folding@home distributed computing project, Weber applied theoretical physics to develop models of nonequilibrium protein dynamics at unprecedented time scales, on the order of milliseconds.
“I basically just used a trajectory-based formulation of statistical mechanics, like some that have been around for many years,” Weber said. “This is a practical implementation of those kinds of ideas, one that allows us to connect path integrals to longer time scale biology.” Focusing on kinases, the primary signaling engines in cells, and GPCRs, targets for a large percentage of prescription drugs, Weber analyzed protein systems in both glassy and heat-dissipating states. By incorporating basic laws of thermodynamics and statistical mechanics, he was able to bias the trajectories of proteins, taking space and time into account to figure out all the possible paths the molecules could take and which of those paths were most important in a given context.
“We estimated a propagator, like one you might find in quantum field theory, that can predict equilibrium protein trajectory probabilities. We then used this propagator to bias trajectories, to see how heat might be dissipated through the system or where the system might be slowed down and trapped,” he said.
Like manually pulling on the pistons of a car’s engine, Weber applied forces to proteins’ moving parts and tracked where they generated heat, enabling him to find out where work was being done and to probe their functional dynamics without having to simulate the full environment of the cell.
“Proteins are the heat engines that drive our bodies’ movements and everything else that happens in living organisms,” Weber said. “We know from thermodynamics that whenever you do meaningful work in an engine, you’re always going to have to dissipate some amount of heat into the environment. We found that yes, we could indeed carefully tug at these models of biomolecules wiggling inside a computer, and then systematically track where heat was being dumped into the environment. It turns out you can correlate these dissipative motions very well with the functions of some proteins.”
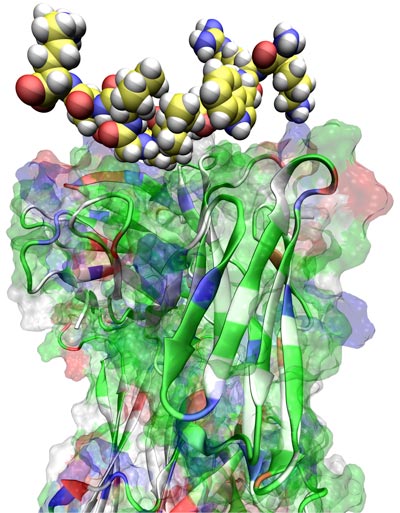
Image right: T cell receptor (TCR; bottom) of an HIV elite controller—a patient whose body controls an HIV infection at a clinically elite level compared to the rest of the population—in complex with the HIV peptide KK10 (top). Elite controllers of HIV exhibit exceptional immune responses upon exposure to the virus, perhaps holding the key to the development of an effective HIV vaccine. Improved recognition of HIV peptides by killer T cells (which have TCRs like that pictured) is thought to be a crucial component of elite controller immunity.
Weber, 28, earned his PhD in chemistry from Stanford in 2014 with the support from his Fannie and John Hertz Foundation Fellowship, and has taken his ideas to the Soft Matter Theory and Simulation group at the IBM Thomas J. Watson Research Center in New York, where he has unfettered access to some of the world’s fastest supercomputers.
A postdoctoral fellow for IBM, Weber has focused his research on the computational design of vaccines for HIV and cancer immunotherapies. Weber’s main thrust is trying to use physical simulations and artificial intelligence to study protein dynamics. He’s looking at how the immune system works, specifically researching why specific T cell receptors in some patients are so effective against HIV peptides and how to mimic these receptors to create working components of a vaccine.
“In terms of HIV, people have been trying to develop vaccines for 30 years, so it’s challenging,” Weber said. “It’s definitely a hard project but one that should be considered worthwhile. We still don’t know everything about how our immune systems work, even on a very basic, phenomenological level.”
While the work is both “rewarding and daunting,” Weber said he’s encouraged by the increasing capabilities of supercomputers and other transformative technologies. Within 10 years, Weber hopes to be able to fully simulate protein-protein interactions on an atomistic level, and to continue using physics to bridge the gap between foundational scientific principles and knowledge of how cells work.
“The ultimate goal would be to develop therapies for diseases—and that’s something we’re trying to do—but it’s also just an amazing physical problem,” Weber said. “Using these very basic theoretical physical principles (to understand cells) is exciting to me and to a lot of other physicists as well.”
Currently spending a lot of time in New York City, Weber said he wants to continue working at the intersection of industry and academia, focusing his attention on biological systems, physical simulations, and machine learning.